





Index: Volume 3 Number 1
Next Article: Oligo Coupling made easy and fast
 |
Abstract
The PCR amplification efficiency using Taq-FORCE™ Amplification System's MIGHTY™ Buffer was compared with a common Tris/KCl PCR buffer system for amplification of DNA templates with and without biotin-14-dCTP incorporation. Amplification of PCR fragments of the mouse IL-1b cDNA and p53 cDNA were used for comparison. MIGHTY™ Buffer performed equally well as the common PCR buffer system for amplification of the PCR fragment using unlabeled-dNTPs. However, use of MIGHTY™ Buffer vs. the common Tris/KCl buffer greatly enhanced the amplification efficiency when biotin-14-dCTP was included in the reaction mix.
Introduction
Non-isotopic detection has become the method of choice for gene identification. This method relies on a non-isotopic labeled nucleic acid probe, for example a biotinylated probe, which hybridizes to the specific gene sequence of interest. Next a reporter enzyme conjugated with a molecule, for example a biotin-specific streptavidin, that specifically binds to the non-isotopic label is introduced. Then an appropriate substrate is added to generate a detectable signal which may be colorimetric (ELISA), chemiluminescent or fluorescent. The greater number of biotinylated moieties in the hybridized probe, the stronger the signal generated. One of the methods for incorporating biotin at multiple sites along the probe sequence is use of biotin-labeled nucleotide analogs during amplification of the probe by PCR. In this study we evaluated the efficiency of biotin-14-dCTP incorporation into two different sequences using PureBiotech LLC's Taq-FORCE™ DNA Amplification System containing MIGHTY™ Buffer and a common Tris/KCl PCR buffer supplied with a major competitor's Taq DNA polymerase.
Methods and Results
Comparison of PCR amplification efficiency was performed as follows: 50 µl reactions were prepared using 10,000 copies of mouse p53 template DNA and corresponding primer set (CLONTECH), 1.5 mM MgCl2, 2.5 U cloned SuperTaq™ DNA Polymerase (5 U/µl) (PureBiotech LLC), 200 µM each dNTPs (PureBiotech LLC), and either MIGHTY™ Buffer (PureBiotech LLC) or the common Tris/KCl PCR buffer supplied by a leading competitor. 35 cycles were performed using PCR conditions optimized for the mouse p53 cDNA. A second set of reactions, designed to incorporate biotin into the PCR amplified sequence, were prepared as above with the addition of 50 µM dCTP + 150 µM biotin-14-dCTP (Life Technologies) to the reaction mix. The 25%:75% ratio of dCTP:biotin-14-dCTP was used per the recommendation of the supplier of the biotin-14-dCTP. 35 cycles of PCR were performed. 2 µl from each reaction were resolved on a 2% agarose (1X TAE) gel and visualized by UV following ethidium bromide staining. MIGHTY™ Buffer (Figure 1, Lane 6) performed equally well as the common Tris/KCl PCR buffer (Figure 1, Lane 5) for amplification of the PCR fragment using unlabeled-dNTPs. However, use of MIGHTY™ Buffer (Figure 1 Lanes 3-4), as compared with the common Tris/KCl PCR buffer (Lanes 1-2), greatly enhanced the amplification efficiency when biotin-14-dCTP was included in the reaction mix.
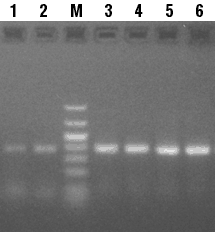 |
Figure 1: Comparison of PCR Amplification Efficiency of Mouse p53 cDNA with Different PCR Buffers. PCR reaction conditions were as described in the text. Lane M, DNA size marker; Lanes 1 and 2, common Tris/KCl buffer in the presence of biotin-14-dCTP; Lanes 3 and 4, MIGHTY™ buffer (PureBiotech LLC) in the presence of biotin-14-dCTP; Lane 5, common Tris/KCl buffer without biotin-14-dCTP; and Lane 6, MIGHTY™ Buffer (PureBiotech LLC) without biotin-14-dCTP. |
Once enhanced amplification efficiency with incorporation of biotin-14- dCTP was established using MIGHTY™ Buffer, a second study was then performed to evaluate the effect of increasing the amount of biotin-14-dCTP in the reaction mix on the intensity of the PCR signal. As expected, the amplification efficiency of the PCR reaction decreased as the ratio of biotin-14-dCTP increased in the reaction mix (Figure 2). When 100% biotin-14-dCTP was used, the amplification was completely diminished (Figure 2, Lane 5). The corresponding increase in molecular weight of the amplified fragment illustrated by the slower movement of the fragment combined with the increase in biotin-14-dCTP concentration (Figure 2, compare Lane 1: 0% biotin-14-dCTP and Lane 4: 75% biotin-14-dCTP) indicates increased incorporation of biotin into the DNA. These results show a dramatic increase in the efficiency of labeled nucleotide incorporation during PCR using MIGHTY™ Buffer.
* PCR is covered by U.S. patents owned by Hoffmann-LaRoche
| Figure 2: MIGHTY™ Buffer (PureBiotech, LLC) PCR amplification of mouse IL-1b cDNA with Increased Ratio of dCTP:Biotin-14-dCTP. Lane M, DNA size marker; Lane 1, 100% dCTP:0% biotin-14-dCTP; Lane 2, 75% dCTP:25% biotin-14-dCTP; Lane 3, 50% dCTP:50% biotin-14-dCTP; Lane 4, 25% dCTP:75% biotin-14-dCTP; Lane 5, 0% dCTP:100% biotin-14-dCTP. | 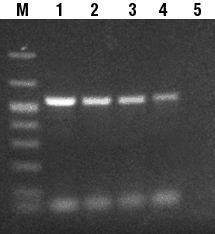 |
Next Article: Oligo Coupling made easy and fast
© 2004-2023 PureBiotech LLC All rights reserved.