





Index: Volume 3 Number 1
Next Article: Improved Labeling Efficiencies in PCR
- True BACKGROUND-FREE Cloning
- Eliminate the colony screening step — all colonies are positive
- Works with any strain of E. coli
- No need for dephosphorylation, complete digestion, or gel purification
- One hour, one step — sure to work every time
Most prokaryotic cloning vectors that are in common use today provide a marker gene (ampicillin, kanamycin, tetracycline, etc...) to select against the growth of cells that do not contain the vector. However, cells will grow if they harbor the vector whether or not they contain the DNA fragment one is attempting to clone. To select the bacteria containing recombinant molecules, a second selection system is needed to discriminate cells with the cloned fragment from cells containing non-recombinant molecules. Traditionally, a type of positive selection system that has seen wide use is the blue-white color selection based on the b-galactosidase enzyme that is available in pUC vectors and their derivatives. In this case, there is no growth selection against a vector without insert so many colonies are formed carrying the vector that has either not been cut in the first place, or re-ligated without the insert. For this screening system, extra measures are then required to increase the percentage of positive colonies via more complete restriction digests and dephosphorylation to prevent re-ligation without insert.
New Generation Cloning
A second generation of positive selection systems is now available. This system is based on the inactivation of a toxic protein by the cloning process. With this new vector, bacterial cells containing the vector without insert are killed by the toxic peptide, and only the cells containing recombinant molecules can grow on the selective media. This process, can lead to 95-100% positive transformants. This is referred to as "background-free" positive selection.
The CloneSure™ pGATA positive selection vector is a 5329 base pair general cloning vector which contains a gene coding for a toxic peptide that inhibits bacterial growth when expression is induced (Figure 1). Ligation of a DNA fragment into the multiple cloning site disrupts the gene coding for the toxic peptide, thus rendering the recombinants non-toxic. The vector, in the absence of the insert, produces a toxic peptide which inhibits bacterial replication. Therefore, only colonies which harbor a plasmid containing your cloned insert will grow. This constitutes a background-free positive selection system for recombinant molecules.
Background-Free Positive Selection
The coding region of toxic peptide in CloneSure™ contains a region of the DNA binding domain of the mouse eukaryotic transcription factor GATA-1. This peptide consists of a zinc finger followed by a region rich in basic amino acids. Trudel et al. (1) have shown that the toxic peptide is able to bind to the bacterial origin of replication, resulting in termination of cell growth. This toxic peptide is expressed as a fusion protein with inactive glutathione-S-transferase protein (GST) which has been disrupted by the in-frame insertion of multiple cloning sites. This fusion protein is under the control of the very strong tac promoter. Expression is induced by the addition of IPTG in the growth medium. To prevent leaky expression of the fusion protein, the CloneSure™ vector also contains the strong lacIq repressor gene. Furthermore, CloneSure™ contains the ampicillin resistance gene and a high copy number origin of replication (colEI type). A T7 RNA polymerase promoter adjacent to the multiple cloning site is designed to allow for the synthesis of RNA transcripts from cloned inserts for use as probes or for in vitro translation. Unlike other similar vectors which require special strains of E. coli to accomplish background-free cloning, CloneSure™ can be used effectively with any strain of E. coli.
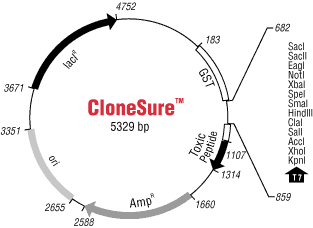 |
Figure 1: Map of the CloneSure™ pGATA Positive Selection Vector |
Save Time
Use of the CloneSure™ vector system eliminates time consuming steps required in standard cloning protocols. Most standard cloning protocols require that the vector termini be dephosphorylated after digestion to prevent re-ligation of the vector. Many protocols also require the vector be completely digested by using a large excess of restriction enzymes and/or by purifying the digested form of the vector. These steps are not only time-consuming, but also reduce cloning efficiency.
Table 1: Comparison of Standard Vector Systems vs. CloneSure™
| Vector System | CloneSure™ | Standard Vectors | ||
| Total Time | 1 hour | 1/2 - 1 day | ||
| Number of Steps | 1 | 4 | ||
| Starting Material (DNA) | 0.5 - 1 µg | 10 - 50 µg | ||
| Positive Clones (recombinants) | 80 - 100% | 5 - 25% |
Performance
To have sufficient vector DNA to perform all ligation experiments and controls, it is common practice to start with 10 to 50 µg of vector DNA. It is therefore difficult to clone DNA fragments available in limited quantities. The use of CloneSure™ positive selection system eliminates the worry about the quality of your vector preparation and the quantity of vector available. DNA from standard, even "dirty" mini-preps can be used directly without further purification because partial digestion is not a problem. Since the only manipulation is to inactivate the restriction enzyme used to cut the vector, loss of starting material is reduced to a minimum, and a cloning experiment can be completed with less than 1µg of starting material. (Table 1)
Applications
Whatever the cloning application is, the CloneSure™ positive selection vector will screen the right clones for you without radioactivity. From the routine cloning of DNA using positive selection to the construction of cDNA libraries, it works for you. In sequencing, the selection system allows directional cloning using the versatility of the Multiple Cloning Site (MCS) without the need to purify the vector or the recombinant fragments. The Ampr/GATA selection ensures the growth of only your recombinants with the additional flexibility of using a variety of common E. coli strains.
Typically, after ligation of 100 ng of CloneSure™ with an insert:vector ratio of 2:1 in a 10 µl reaction volume, transformation of DH5 competent cells (efficiency of 3 x 107 transformants/µg DNA) yields more than 104 colonies and greater than 90% recombinants (Figure 2). CloneSure™ is supplied at 0.5 µg/µl and each vial contains enough vector for 100 ligations.
1. Trudel, P., Provost, B., Massie, B., Chartrand, and Wall, L. (1996) BioTechniques 20, 684-693.
Next Article: Improved Labeling Efficiencies in PCR
© 2004-2023 PureBiotech LLC All rights reserved.